The challenge
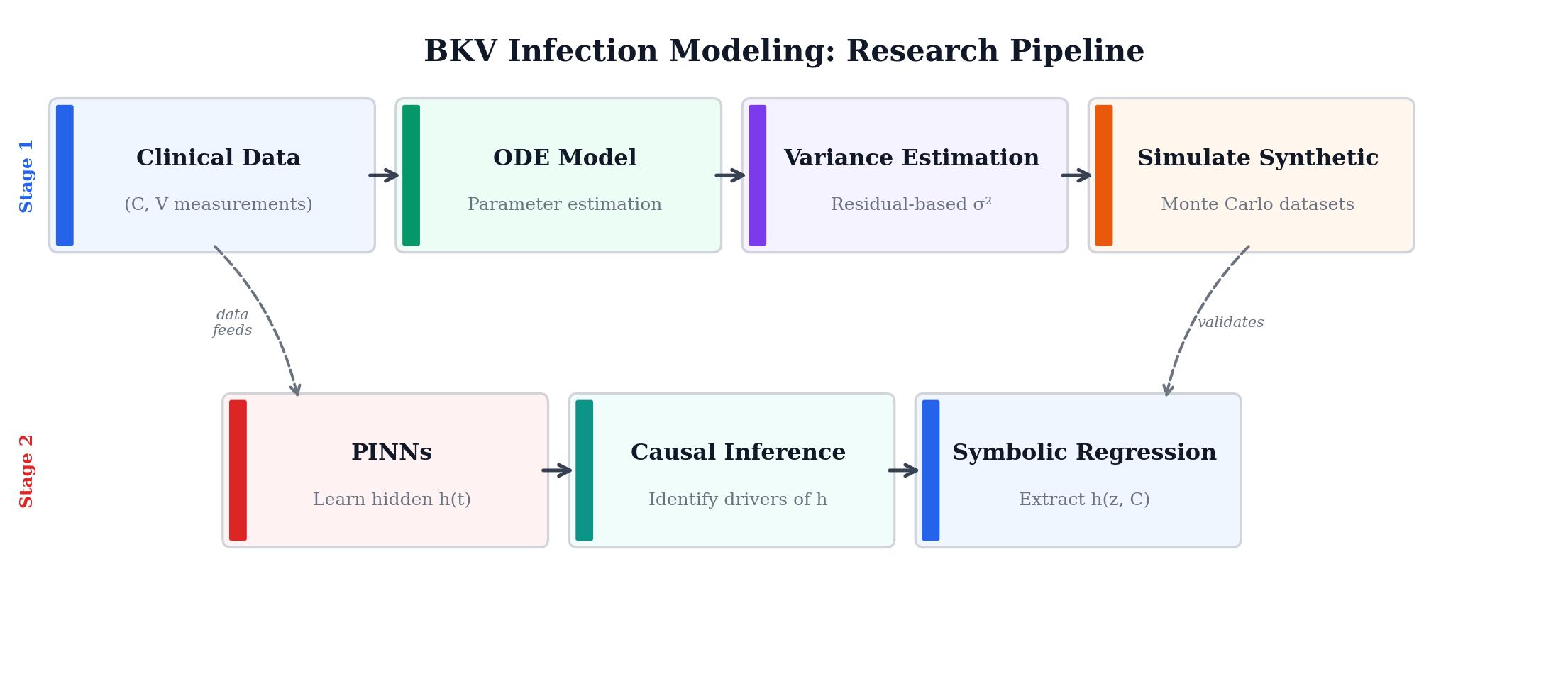
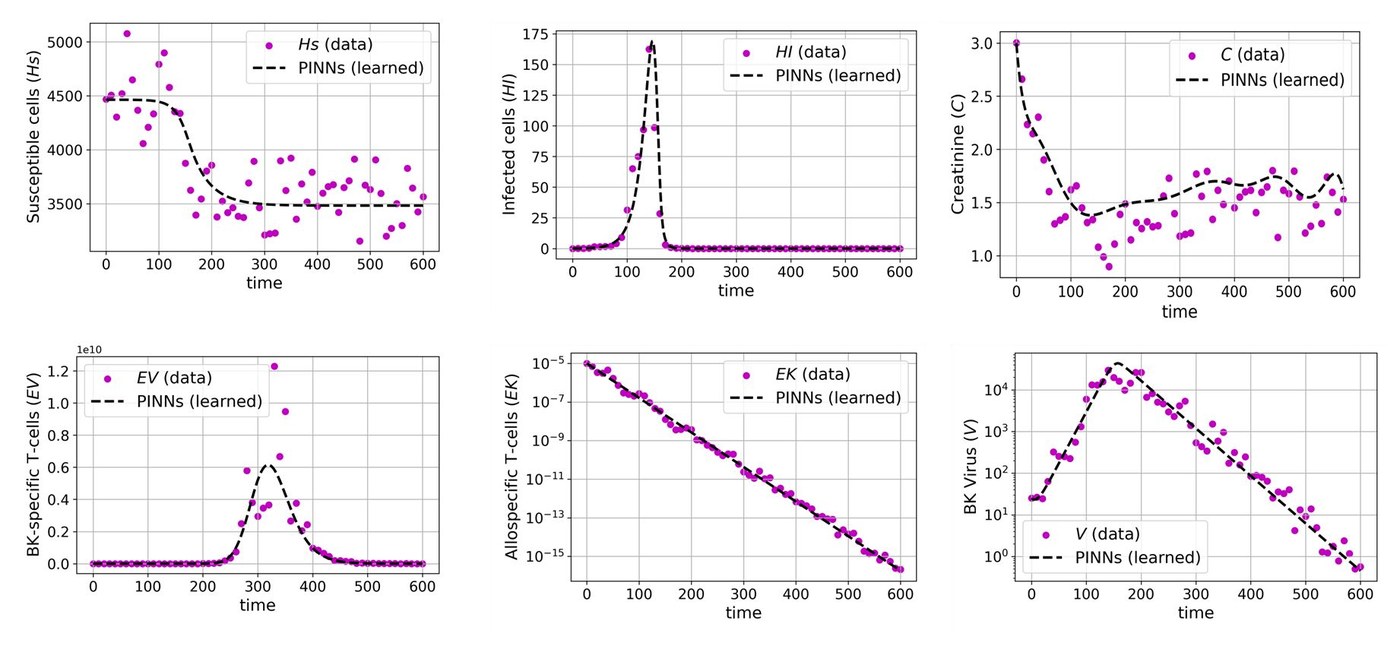
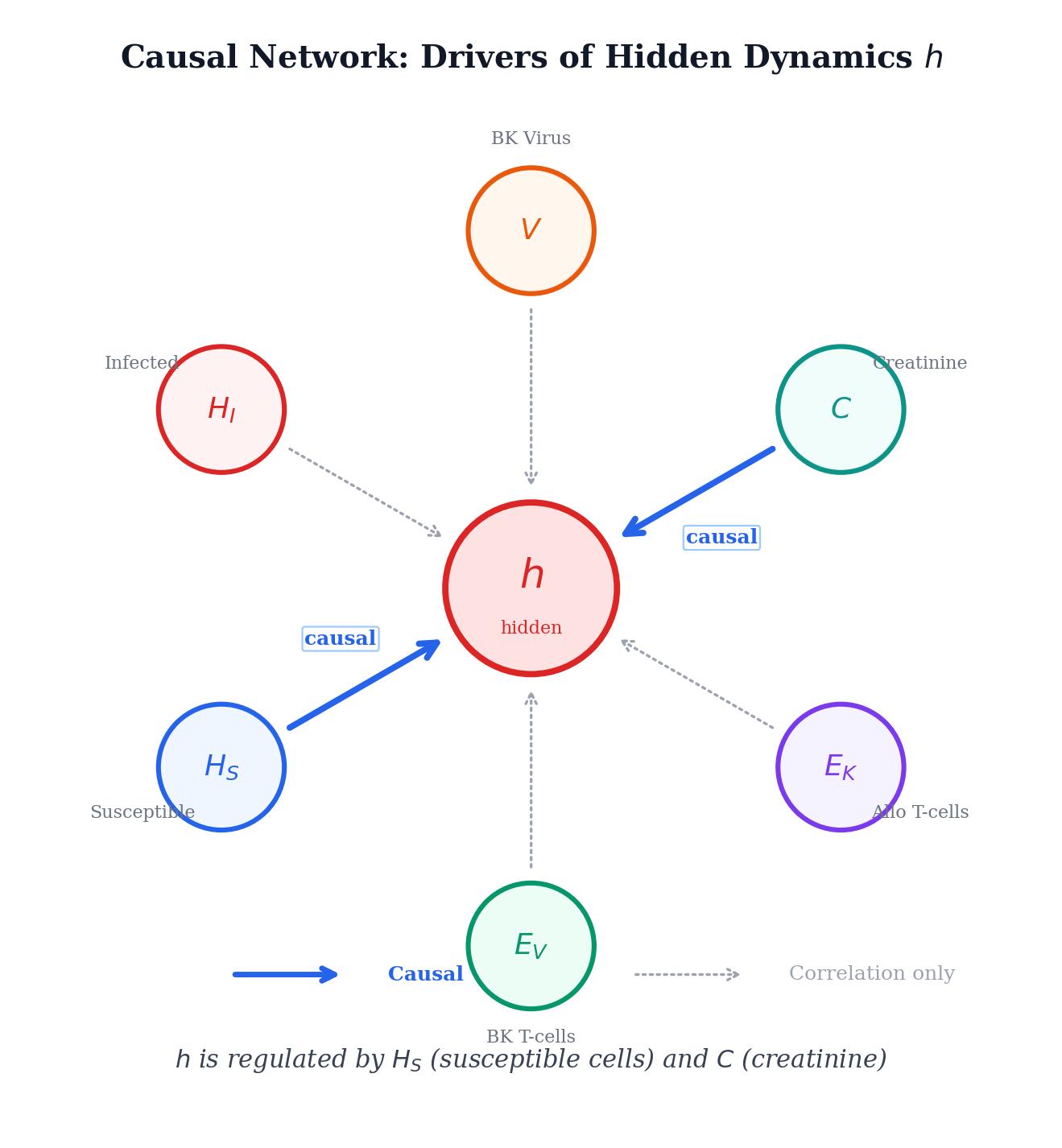
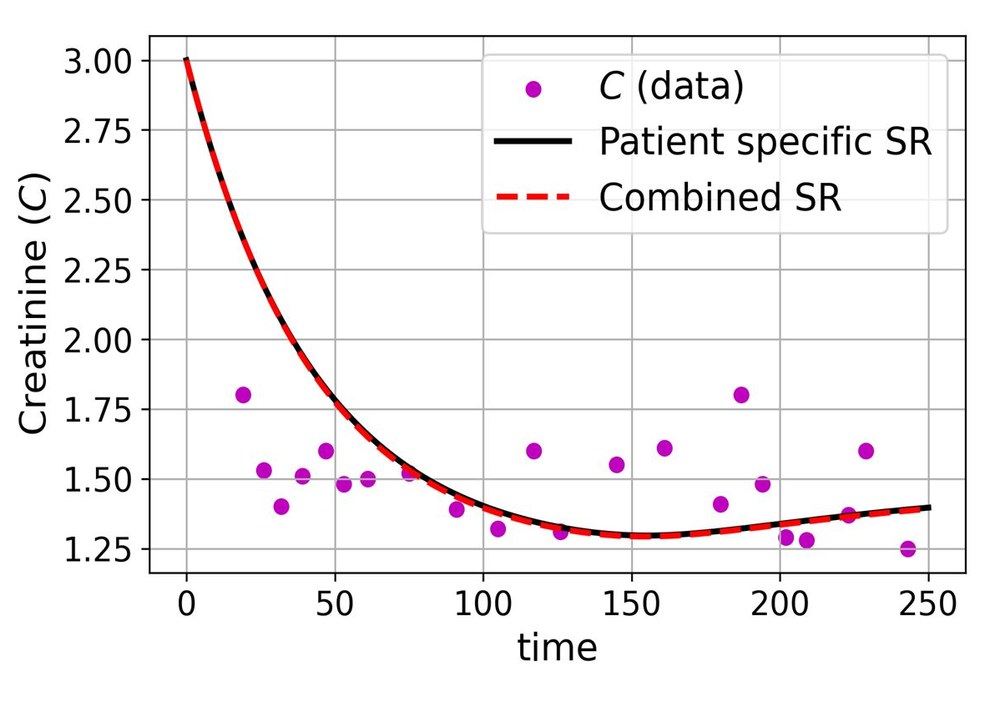
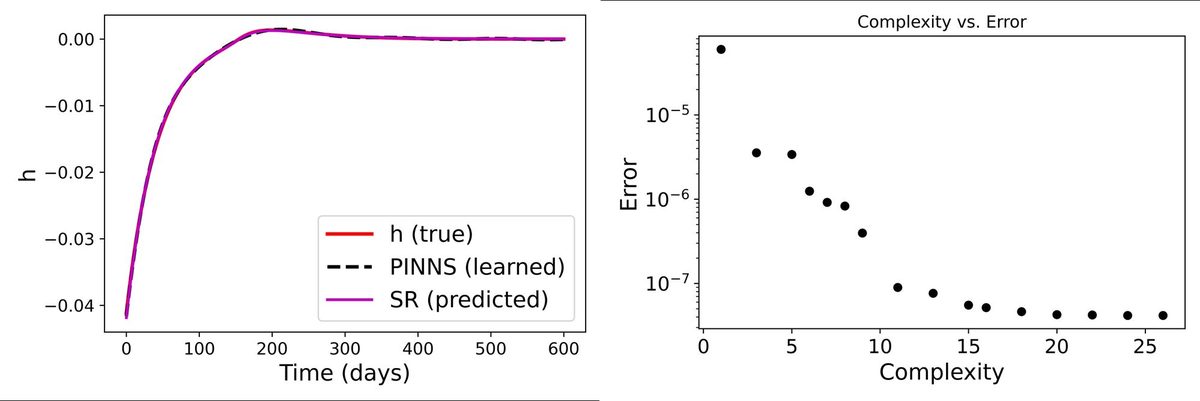
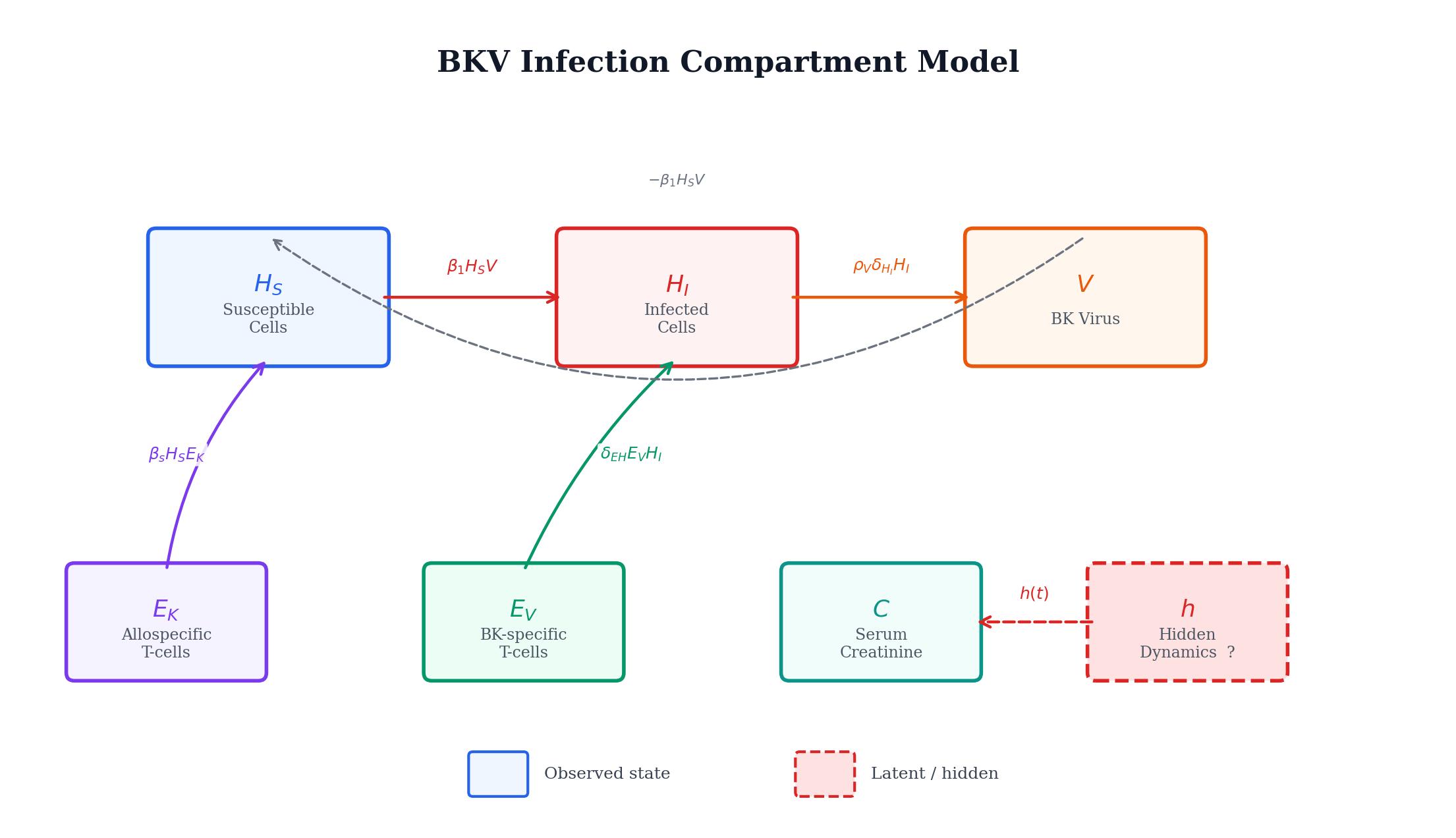
We have a mathematical model that tracks six biological compartments: susceptible cells, infected cells, BK virus, two T-cell populations, and serum creatinine. The model works well for most compartments, but the creatinine equation has a gap. A function \(h(t;\theta)\) on the right-hand side captures unknown dynamics we don't have a mechanistic theory for. The question is: can we learn what \(h\) should look like directly from patient data, without guessing its form in advance?